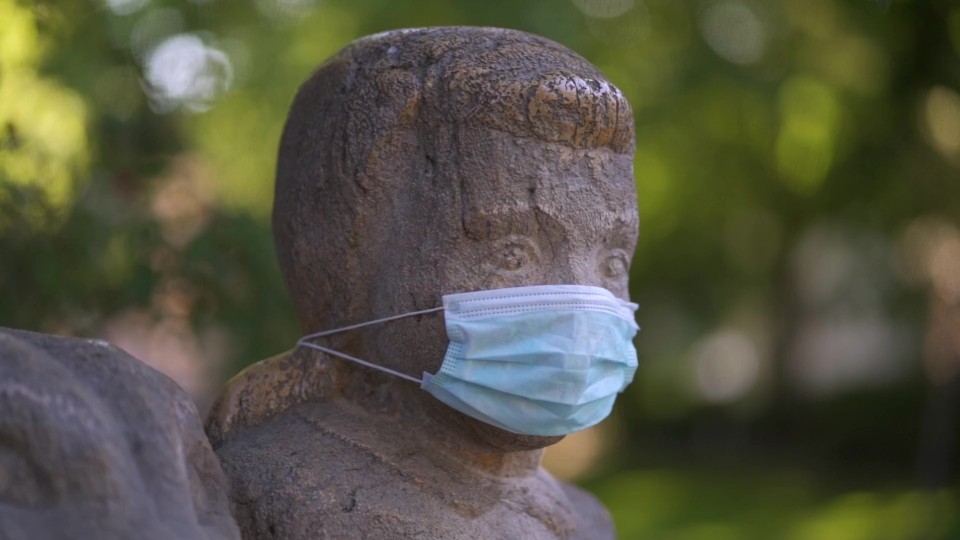
From what I understand, it's not consistent at all. That's why you have talk of "variants." We've heard about the more famous ones (delta and omicron), but according to Andrew Kaufman, there are well over 5.5 Million "variants," or variations of the genetic sequence.
View: https://www.bitchute.com/video/3afWsXJUybnh/
Yes, in addition to this I think these are worth reading, @LLight
"“Reproducibility of computational genomic studies has been considered as a major issue in recent times. In this context, we have characterised workflows on the basis of approach used for their definition and implementation. To evaluate reproducibility and provenance requirements, we implemented a complex variant discovery workflow using three exemplar workflow definition approaches. We identified numerous implicit assumptions interpreted through the practical execution of the work-flow, leading to recommendations for reproducibility and provenance, as shown in Table 1.”
“In the field of genomic data science, accuracy and reproducibility remains a considerable challenge due to the sheer size, complexity, and dynamic nature plus relative inventiveness of the quantitative biology approaches. The accuracy and reproducibility challenge does not just block the path to new scientific discoveries, more importantly, it may lead to a scenario where critical findings used for medical decision making are found to be incorrect (Huang and Gottardo, 2013). NPARS has been developed to meet the unmet need of improving accuracy and reproducibility in genomic data science. Currently, a limitation of our system is the requirement of the user to put their data into a standardized format for import into NPARS. These steps are not automated.”

The Reproducibility Crisis in Genomics
It has been known since at least 2005 that much of the scientific literature being published is fundamentally flawed, non-reproducible, and/or outright fraudulent. Relating to the (pseudo)science o…
 viroliegy.com
viroliegy.com
The article below is especially useful to read on this topic:
"...observed viral genomes often deviate considerably from reference genomes demanding use of exhaustive alignment approaches;"
"Various software tools have been developed to accommodate the unique challenges and use cases associated with characterizing viral sequences; however, the quality of these tools varies, and their use often necessitates computing expertise or access to powerful computers, thus limiting their usefulness to many researchers."

“Viral” Genomics: Nothing but Strings of Letters in a Data Bank
“Although all that is terrific, says Calisher, a string of DNA letters in a data bank tells little or nothing about how a virus multiplies, which animals carry it, how it makes people sic…
 viroliegy.com
viroliegy.com